Applying genetic information
Identifying risks to fishes and sharks, including threatened and endangered species can be challenging in the vast marine environment. Understanding the diversity, connectivity patterns, breeding areas of marine fishes and the environmental factors that drive them is critical for the sustainable management of fisheries. One of the tools we use to understand fish diversity and connectivity is DNA.
DNA is the genetic code for all living organisms.
By analysing DNA from fish samples, we can gain a deeper understanding of:
- taxonomic biodiversity
- population structure and genetic connectivity of marine fish species.
Harnessing molecular genetics and DNA barcoding
Rapid developments in molecular techniques suitable for examining biodiversity are helping to unlock valuable genetic information from our biological collections.
Molecular genetics and DNA analyses are extremely powerful, because DNA:
- is an information rich molecule found in all living cells
- provides identification of individuals, populations, or species of fishes
- is inherited, therefore genetic lineages and relatedness of fish can be traced
- can be analysed at any life cycle stage.
DNA - and more importantly RNA - degrades after death, so tissue samples need to be collected and stored appropriately.
We accurately assign specimens to species with the help of methods, such as sequencing specific areas of the DNA from individual fish (e.g. COI barcoding) and next generation metabarcoding (which simultaneously identifies multiple specimens in a sample (e.g. fish eggs or larvae)). This is done alongside alpha taxonomy (which uses morphology and meristics).
We also use DNA markers such as microsatellites, single nucleotide polymorphisms and sequencing of genes/large sections of the genome, for the assessment of genetic diversity, familial relationships and connectivity among individuals, populations, and species.
DNA barcoding analysis
We are one of the largest and ongoing contributors of fish sequences to the international Barcode of Life initiative (BOLD), and as part of the BioPlatforms Australia DNA barcoding initiative, we are developing framework datasets for Australian fishes.
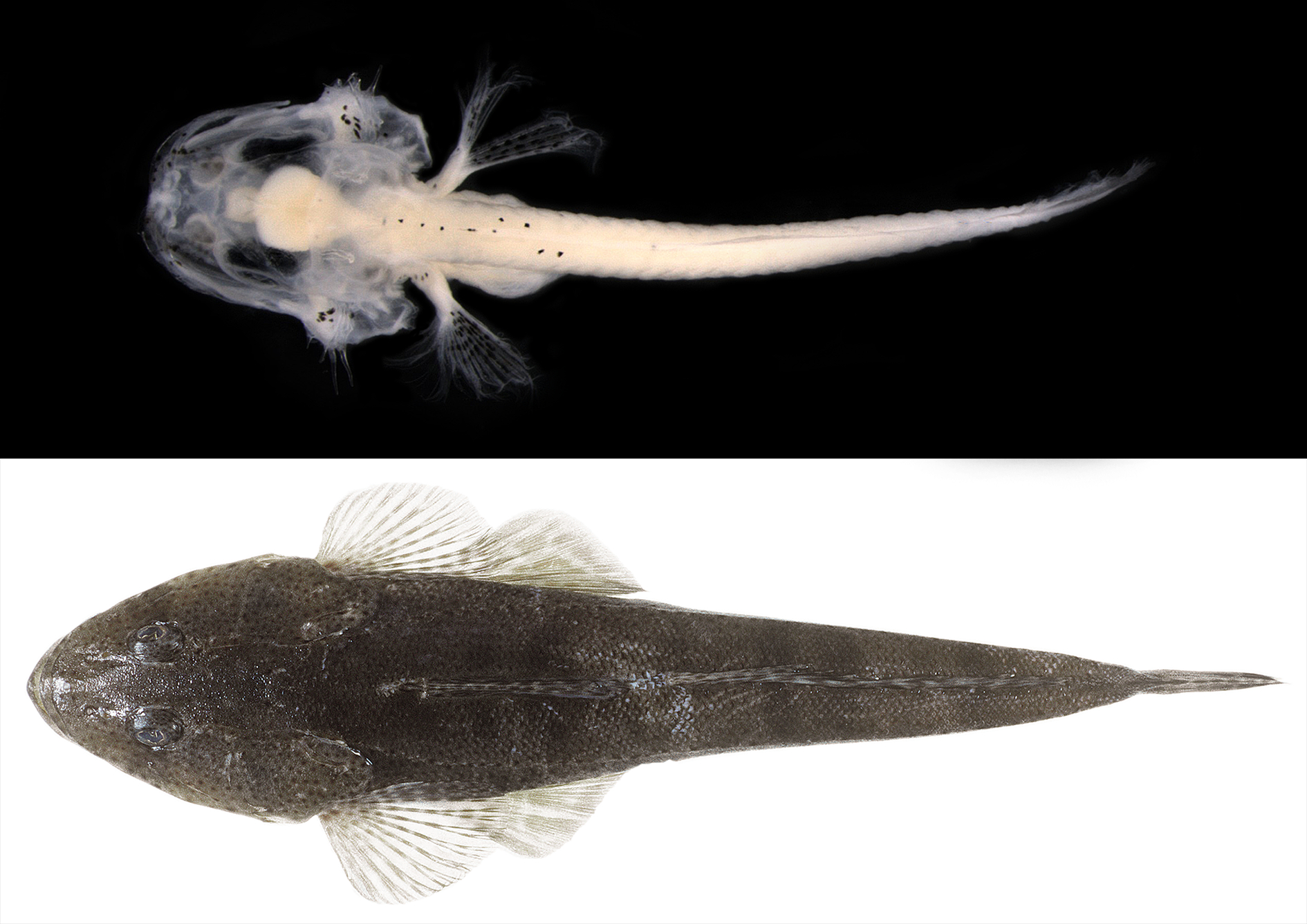

Through the assembly of a standardised reference library of fish DNA barcodes, researchers provide important genetic information for both the research community and industry via:
- accurately assigning specimens (eggs to adults, tissue fragments, prey, stomach contents; including from formalin fixed specimens) to species
- linking Australian larval fishes to their adult forms through morphology and DNA barcoding
- undertaking rapid taxonomic surveillance of market or unknown samples (e.g. from shark and ray egg cases)
- advancing the genetic and ecological understanding of fish (which leads to more sustainable marine fish management)
- increasing our knowledge of the biodiversity of Australian fishes.
Molecular phylogenetics and connectivity of fish
We undertake molecular phylogenetic studies of key fish groups (i.e. wrasse and flatheads), which improve our understanding of historical evolutionary processes in Australian waters.
Our partners include the National Environmental Science Programme, the Australian Centre for International Agricultural Research and Tree of Life Chondrichthyes project.
Additionally, using DNA from the ANFC, alongside material from other collections, researchers are:
- developing a greater understanding of the genetic connectivity of marine fish species in domestic and international waters, (this is important for the sustainable management of fisheries stocks and for conservation of endangered species (e.g. spotted handfish, Swan galaxias, silver tip, bull and hammerhead sharks))
- identifying risks to threatened and endangered species.
Environomics Future Science Platform
Our research on fish collection genomics (including degraded DNA studies) is part of the Environomics Future Science Platform research.
Applying genetic information
Identifying risks to fishes and sharks, including threatened and endangered species can be challenging in the vast marine environment. Understanding the diversity, connectivity patterns, breeding areas of marine fishes and the environmental factors that drive them is critical for the sustainable management of fisheries. One of the tools we use to understand fish diversity and connectivity is DNA.
DNA is the genetic code for all living organisms.
By analysing DNA from fish samples, we can gain a deeper understanding of:
- taxonomic biodiversity
- population structure and genetic connectivity of marine fish species.
Harnessing molecular genetics and DNA barcoding
Rapid developments in molecular techniques suitable for examining biodiversity are helping to unlock valuable genetic information from our biological collections.
Molecular genetics and DNA analyses are extremely powerful, because DNA:
- is an information rich molecule found in all living cells
- provides identification of individuals, populations, or species of fishes
- is inherited, therefore genetic lineages and relatedness of fish can be traced
- can be analysed at any life cycle stage.
DNA - and more importantly RNA - degrades after death, so tissue samples need to be collected and stored appropriately.
We accurately assign specimens to species with the help of methods, such as sequencing specific areas of the DNA from individual fish (e.g. COI barcoding) and next generation metabarcoding (which simultaneously identifies multiple specimens in a sample (e.g. fish eggs or larvae)). This is done alongside alpha taxonomy (which uses morphology and meristics).
We also use DNA markers such as microsatellites, single nucleotide polymorphisms and sequencing of genes/large sections of the genome, for the assessment of genetic diversity, familial relationships and connectivity among individuals, populations, and species.
DNA barcoding analysis
We are one of the largest and ongoing contributors of fish sequences to the international Barcode of Life initiative (BOLD), and as part of the BioPlatforms Australia DNA barcoding initiative, we are developing framework datasets for Australian fishes.
Through the assembly of a standardised reference library of fish DNA barcodes, researchers provide important genetic information for both the research community and industry via:
- accurately assigning specimens (eggs to adults, tissue fragments, prey, stomach contents; including from formalin fixed specimens) to species
- linking Australian larval fishes to their adult forms through morphology and DNA barcoding
- undertaking rapid taxonomic surveillance of market or unknown samples (e.g. from shark and ray egg cases)
- advancing the genetic and ecological understanding of fish (which leads to more sustainable marine fish management)
- increasing our knowledge of the biodiversity of Australian fishes.
Molecular phylogenetics and connectivity of fish
We undertake molecular phylogenetic studies of key fish groups (i.e. wrasse and flatheads), which improve our understanding of historical evolutionary processes in Australian waters.
Our partners include the National Environmental Science Programme, the Australian Centre for International Agricultural Research and Tree of Life Chondrichthyes project.
Additionally, using DNA from the ANFC, alongside material from other collections, researchers are:
- developing a greater understanding of the genetic connectivity of marine fish species in domestic and international waters, (this is important for the sustainable management of fisheries stocks and for conservation of endangered species (e.g. spotted handfish, Swan galaxias, silver tip, bull and hammerhead sharks))
- identifying risks to threatened and endangered species.
Environomics Future Science Platform
Our research on fish collection genomics (including degraded DNA studies) is part of the Environomics Future Science Platform research.
